I have recently contributed to the characterization of a new human mRNA export factor called Chtop for Chromatin target of Prmt1 (also termed SRAG, Fop, or c1ORF77). In addition to its known role in transcription 28, our study has revealed that Chtop is a new component of the THO-TREX complex and is an mRNA export co-adaptor. Namely, it is associated with core THO-TREX components in vivo and it binds to the central NTF2-like (NTF2L) domain of the essential mRNA export receptor Nxf1 to form a trimeric complex Chtop:Nxf1:p15. Its binding to Nxf1 is promoted by the adaptor Alyref (also called Aly or REF) and, as a consequence, the RNA binding activity of Nxf1 and the process of mRNA export itself are stimulated.
In this respect, Chtop mode of action resembles that of the co-adaptor protein Thoc5 described previously 24 and in the schematics above. However, our work also shows that the two proteins differ in that Chtop requires methylation of its numerous arginines to bind Nxf1:p15. While this post-translational modification has no effect on Chtop interaction with the RNA helicase Uap56, it prevents Chtop from interacting with Alyref. Finally, some of our results suggest that another consequence of the methylation of Chtop arginines is a decrease of its RNA binding activity. Interestingly, the Wilson lab has previously shown that the RNA binding activity of Alyref is also decreased by arginine methylation and this modification enhances the mRNA handover process 29. Therefore, post-translational arginine methylation appears to promote mRNA export in human cells by stimulating the transfer of the mRNA to the export receptor Nxf1:p15 by two distinct mechanisms:
· decrease of the RNA binding activities of the adaptor Alyref and co-adaptor Chtop,
· stimulation of Chtop binding to Nxf1:p15.
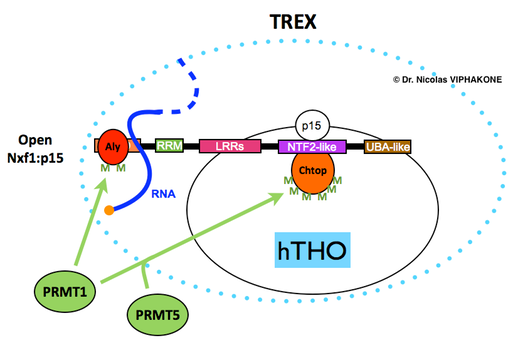
Chtop is a component of the human THO-TREX complex and is regulated by arginine methylation. Arginine methylation of Chtop by PRMT5 seems to prime its arginine methylation by PRMT1 28. Upon methylation, Chtop is able to bind the NTF2L domain of the essential mRNA export receptor Nxf1. In conjunction with the binding of methylated Aly (a.k.a Alyref), methylated Chtop stimulates transfer of the mRNA to Nxf1:p15 and thereby promotes mRNA export in human cells. The green "M" stand for methylated arginines.
In addition to performing various RNA binding and protein-protein interaction assays for this study, I have designed an experiment which has shown the presence of both unmethylated and methylated forms of Chtop in the nucleus. The unmethylated form was only detected in this cellular compartment, which suggests that Chtop could potentially interact with Alyref in the nucleus at an early stage of mRNP formation. This Chtop:Alyref interaction, the subsequent binding of methylated Chtop to Nxf1, and the methylation of Alyref are therefore very likely to be tightly and temporally co-regulated during mRNP formation to allow for efficient mRNA transfer to Nxf1:p15. Several facts suggest that this must be the case:
· Alyref is a substrate for the PRMT1 enzyme 29 and one of the initial names of Chtop was Fop (Friend of PRMT1) due to its interaction with this enzyme 28.
· In yeast, Yra1 and Hmt1p (orthologues of Alyref and PRMT1 respectively) are recruited during transcription alongside with the RNAPII-dependent transcription machinery 18,30,31.
· Very recently, Chtop (a.k.a Srag, Fop, c1orf77) has been found associated with PRMT1 and with the 5FMC complex to activate the RNAPII-dependent transcription of some genes 32.
Therefore post-translational arginine methylation of mRNA export factors appears to be important to finely regulate RNA-protein and protein-protein interactions in close connection to transcription in order to ultimately achieve timely transfer of the mRNA to the export receptor Nxf1:p15.
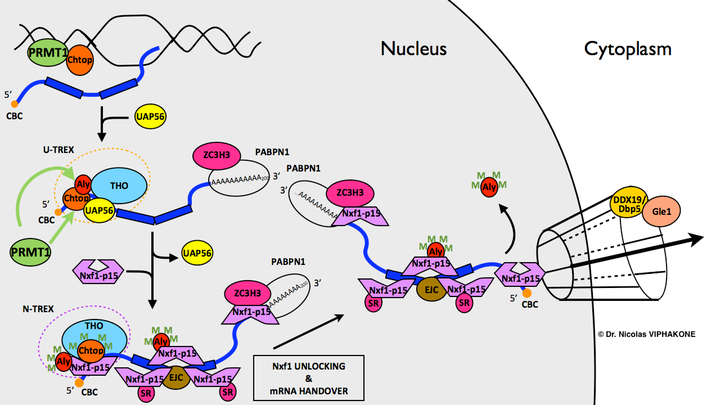
Chtop is an mRNA export co-adaptor that works in conjunction with the adaptor Alyref to promote mRNA export in human cells. Process as known in 2013. Chtop is strongly associated with chromatin where it interacts with PRMT1 32. Loading of Chtop onto the mRNA requires the RNA helicase Uap56 (a.k.a. Ddx39b or Bat1). Upon arginine methylation by PRMT1, methylated Alyref 29 and methylated Chtop bind Nxf1:p15, stimulate its RNA binding activity, and thereby promote mRNA export. Considering that binding of Thoc5 (see above) and of Chtop on the NTF2L domain of Nxf1:p15 are mutually exclusive but that they can be found in the same complex in vivo, some rearrangements are likely to take place within the THO-TREX complex during mRNP biogenesis. The green "M" stand for methylated arginines. Here, I have designated "U-TREX" the TREX complex containing the RNA helicase Uap56 and "N-TREX" the subsequent form of the TREX complex in which Nxf1 is present and Uap56 is absent.
During the revision process of this Chtop article, we were asked by one of the referees whether the Uap56 binding motif (UBM, Uap56 is also known as Ddx39b or Bat1) was important for Chtop's function in vivo. Given that depletion of Alyref or Chtop by RNAi only leads to a mild block of mRNA export, this led us to test whether a mutant Chtop lacking that UBM would rescue the lethal mRNA export defect observed in a context where both Chtop and Alyref are depleted.
While this kind of question could in principle be easily answered in yeast thanks to the powerful genetic tools available in this organism, addressing this in human cells would be trickier. Indeed, one always faces issues inherent to traditional transfection-based methods: heterogeneity of the population of transfected cells and overexpression-associated phenotypes that can be sometimes difficult to interpret.
Therefore, for this functional analysis of Chtop, we took advantage of a complementation system that I had conceived recently. In the current Flp-In-293/Block-iT system from Invitrogen, a tetracycline-inducible promoter drives the expression of an mRNA encoding EmGFP and pri-miRNAs located in its 3' UTR. Since 2009, I thought that if this construct can express EmGFP, it could as well express another protein, possibly fused to EmGFP for cellular localization and tagging purposes. But it is only very recently that I took the time to try this out for my Luzp4 project and for this Chtop EMBO J. article 33. To this end, I have essentially modified the Block-iT vector by inserting restriction sites and an eGFP peptide linker from pEGFPN1 upstream of the EmGFP sequence. I have then cloned in RNAi-resistant versions of the ORFs of interest and transferred the entire construct into the Flp-In system vector pcDNA5-FRT. As a result, endogenous genes are silenced while “at the same time” expressing either a compensatory cDNA or a mutant version of the endogenous cDNA to get more functional data in vivo.
A new compleme
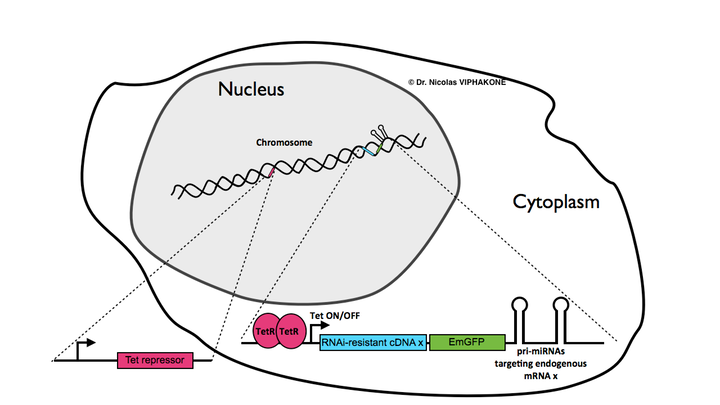
tation system for human cells-based studies
. I have modified the Invitrogen block-iT system in order to express RNAi-resistant cDNAs fused to EmGFP, while targeting the endogenous mRNA of the same gene or of another gene. Using this modified system in combination with the Flp-In-293 cell line, I have performed successful functional complementation studies in homogenous and clonal population of human cells.
Thanks to this system, I have been able to show that the Uap56-interacting motif of Chtop is required for its function in mRNA export in vivo and is essential for cell survival in a context where Alyref is depleted by RNAi. So far, the system has worked brilliantly for this study and this other one and provides the ease and richness of a yeast-like complementation system for human cell-based studies.
References used:
Preceding references: 1-5, 6-16, 17-27.
28. van Dijk, T. B. et al. Friend of Prmt1, a novel chromatin target of protein arginine methyltransferases. Molecular and Cellular Biology 30, 260–272 (2010).
29. Hung, M. L., Hautbergue, G. M., Snijders, A. P. L., Dickman, M. J. & Wilson, S. A. Arginine methylation of REF/ALY promotes efficient handover of mRNA to TAP/NXF1. Nucleic Acids Res. 38, 3351–3361 (2010).
30. Lei, E. P. Intron status and 3'-end formation control cotranscriptional export of mRNA. Genes & Development 16, 2761–2766 (2002).
31. Yu, M. C. et al. Arginine methyltransferase affects interactions and recruitment of mRNA processing and export factors. Genes & Development 18, 2024–2035 (2004).
32. Fanis, P. et al. Five Friends of Methylated Chtop, a complex linking arginine methylation to desumoylation. Mol. Cell Proteomics 11, 1263–1273 (2012).
33. Chang, C-T, Hautbergue, G. M., Walsh, M. J., Viphakone, N., van Dijk, T. B., Philipsen S., and Wilson, S. A. Chtop is a component of the dynamic TREX mRNA export complex. EMBO J 32, 473–486 (2013).