As described above, the pre-mRNA is synthesised in the nucleus of the eukaryotic cell from the DNA template and matured in the same compartment through a series of biochemical modifications: 5' capping, splicing, and 3'-end processing.
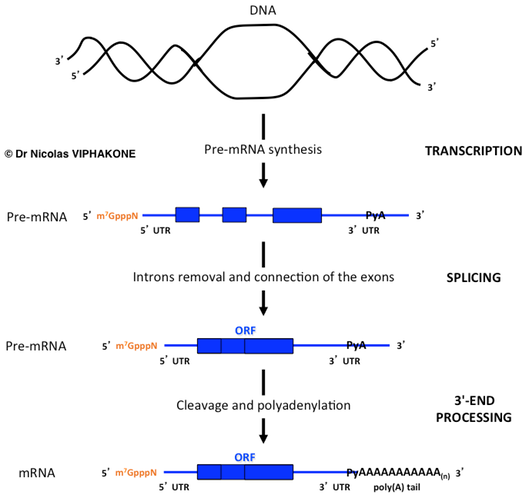
Synthesis and maturation of the pre-mRNA. The capping step corresponds to the addition of m7GpppN (orange) to the 5' end of the pre-mRNA and occurs early in the synthesis stage. Likewise, the poly(A) tail is also not encoded in the DNA template. PyA: Pyrimidine-Adenosine (i.e. CA or UA) is where the cleavage reaction occurs.
During my Ph.D., I worked on pre-mRNA 3’-end processing which consists in two coupled reactions: a site-directed endonucleolytic cleavage in the 3' untranslated region of the RNA, and the addition of a poly(A) tail of a certain length, a process also termed polyadenylation.
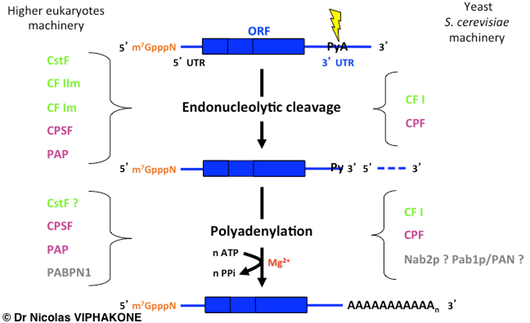
Main steps of pre-mRNA 3'-end processing. A site-directed endonucleolytic cleavage (lightning) takes place in the 3’ untranslated region (3' UTR) of the pre-mRNA and generates two fragments. The 3' fragment is rapidly degraded (dashed line). The 5' fragment is subjected to a polyadenylation reaction which results in the addition of a poly(A) tail to the mRNA. The cellular factors (individual proteins or proteic complexes) performing the pre-mRNA 3'-end processing are indicated on the sides. Functionally equivalent factors are colored similarly. The question marks indicate some uncertainties regarding the involvement of the factors in the corresponding reaction. ORF: Open Reading Frame.
I have thoroughly studied the poly(A)-binding hnRNP Nab2p and its involvement in mRNA polyadenylation regulation in yeast. I have shown that Nab2p uses its CCCH zinc finger region in conjunction to its RGG-box to define and protect the size of the poly(A) tail added to the mRNA in vitro 1. I have also been involved in other studies related to mRNA 3'-end formation:
· another study showing the role of Clp1p, a subunit of the CF IA complex, in the assembly and function of the pre-mRNA cleavage and polyadenylation complexes and their recruitment to genes 3.
· functional study of the hnRNP Nab4p/Hrp1p/CF IB in 3'-end processing in vitro.
Since the end of my Ph.D., some proteins homologous to Nab2p have been discovered in human cells by other research groups and have been associated with some neuronal disorders 4,5.
References used:
1. Viphakone, N., Voisinet-Hakil, F. & Minvielle-Sebastia, L. Molecular dissection of mRNA poly(A) tail length control in yeast. Nucleic Acids Res. 36, 2418–2433 (2008).
2. Dheur, S., Nykamp, K. R., Viphakone, N., Swanson, M. S. & Minvielle-Sebastia, L. Yeast mRNA Poly(A) tail length control can be reconstituted in vitro in the absence of Pab1p-dependent Poly(A) nuclease activity. J. Biol. Chem. 280, 24532–24538 (2005).
3. Haddad, R. Maurice, F., Viphakone, N., Voisinet-Hakil, F., Fribourg, S. & Minvielle-Sebastia, L. An essential role for Clp1 in assembly of polyadenylation complex CF IA and Pol II transcription termination. Nucleic Acids Res. (2011).doi:10.1093/nar/gkr800
4. Leung, S. W. et al. Splice variants of the human ZC3H14 gene generate multiple isoforms of a zinc finger polyadenosine RNA binding protein. Gene 439, 71–78 (2009).
5. Kelly, S., Pak, C., Garshasbi, M., Kuss, A., Corbett, A.H. and Moberg, K. New kid on the ID block: Neural functions of the Nab2/ZC3H14 class of Cys3His tandem zinc-finger polyadenosine RNA binding proteins. RNA Biology (2012).